Download additional information
or use tools
associated with our publications
All under the GPL v3 license.

DenovoFusion
Xinwei Zhao, Marc Hotfilder, Eberhard Korsching.
A de novo assembly based fusion gene detection concept based on DNA-seq data of 100 Ewing sarcoma cases
Brief Bioinform. 2026 Jan 7;27(1):bbag059.
Resources :
The study data can be found here:
EGAS00001000855.
The artificial simulation data (10x to 80x) and the Python script are available :
locally.
MAF
Xinwei Zhao, Eberhard Korsching.
Evaluating the Effectiveness of Various Small RNA Alignment Techniques in Transcriptomic Analysis by Examining Different Sources of Variability Through a Multi-Alignment Approach
MDPI Methods Protoc. 2025, 8(3), 65.
Supplementary Information
Resources (84 GB) :
MAF.zip /
SourceForge.
mRNA:
111HUmte,
mte.gtf,
mte.trans.fa.
miRNA:
111HUpmm,
111HUmt,
bt2_mt,
bt2_pir,
bt2_111HU,
bb_mt,
111.mirbase.gtf,
trans.111.mirbase.fa,
trans.111.primary.fa,
m2_miRtRNA.fasta,
mt.miRtRNA.gtf,
mt.trans.fa.
test data:
mRNA.fastq,
miRNA.fastq.
programs:
install.zip.
Eberhard Korsching, Julian Matschke, Marc Hotfilder.
Splice variants denote differences between a cancer stem cell side population of EWSR1‑ERG‑based Ewing sarcoma cells, its main population and EWSR1‑FLI‑based cells.
International Journal of Molecular Medicine 49, no. 3 (2022): 39.
The article is accompanied by
Supplementary_File_1 (0.6 MB).
Further_Information_S1_S2_S3_S4 (13 MB).
Supporting_R_workspace (86 MB).
Sedigheh Gharbi, Zahra Mohammadi, Maryam Saedi Dezaki, Sadat Dokanehiifard, Shahriar Dabiri, Eberhard Korsching.
Characterization of the first microRNA in human CDH1 that affects cell cycle and apoptosis and indicates breast cancers progression.
The article is accompanied by
Supplementary_File (1.3 MB).
Marc Hotfilder, Nikhil Mallela, Jochen Seggewiss, Uta Dirksen, Eberhard Korsching.
Defining a Characteristic Gene Expression Set Responsible for Cancer Stem Cell Like Features in a sub-population of Ewing Sarcoma Cells CADO-ES1.
Int. J. Mol. Sci. 2018, 19, 3908.
Expression data is submitted to ArrayExpress
Supporting information (1.2 MB)
accompanying the article.
Sedigheh Gharbi, Shahriar Khateri, Mohammad Reza Soroush, Mehdi Shamsara, Parisa Naeli, Ali Najafi, Eberhard Korsching*, Seyed Javad Mowla*.
MicroRNA expression in serum samples of sulfur mustard veterans as a diagnostic gateway to improve care.
PLoS One. 2018 Mar 22;13(3):e0194530.
qPCR data is submitted to GEO.
Supporting information (1 MB).
Christiane Schaefer*, Nikhil Mallela*, Jochen Seggewiss, Birgit Lechtape, Heymut Omran, Uta Dirksen, Eberhard Korsching, Jenny Potratz.
Target Discovery Screens Using Pooled shRNA Libraries and Next Generation Sequencing: a Model Workflow and Analytical Algorithm.
PLoS One. 2018 Jan 31;13(1):e0191570.
Use the ProFED
web application and search for profile patterns in experimental designs.
Alternative: local download.
Boecker F, Buerger H, Mallela N,
Korsching E.
TMAinspiration: Decode interdependencies in multi-factorial tissue microarray data.
Cancer Informatics. 2016;15:143–149.
Programs and more: supplementary download.
TMAinspiration was listed in OmicTools from January 19, 2017 to 2020, when the
omicX
project led by Vincent Henry et al. has ended and the website has shut down.
Poos K, Smida J, Maugg D, Eckstein G, Baumhoer D, Nathrath M, Korsching E.
Genomic heterogeneity of osteosarcoma - shift from single candidates to functional modules.
PLoS One. 2015 Apr 7;10(4):e0123082.
Download
OS_network_data.cys (0.4 MB)
R scripts (25 KB)
RMA normalized expression data (8.6 MB).
Poos K, Smida J, Nathrath M, Maugg D, Baumhoer D, Neumann A, Korsching E.
Structuring osteosarcoma knowledge: an osteosarcoma-gene association database based on literature mining and manual annotation.
Poos K, Smida J, Nathrath M, Maugg D, Baumhoer D, Korsching E.
How MicroRNA and Transcription Factor Co-regulatory Networks Affect Osteosarcoma Cell Proliferation.
PLoS Comput Biol. 2013 Aug;9(8):e1003210.
View the supplementary networks:
The Metabolic (C1) or the
Signaling (C2)
microRNA and TF co-regulatory networks
or download the modules:
Met_cluster_1.cys (0.6 MB)
Sig_cluster_2.cys (1.2 MB)
You might want to use Cytoscape.
Buerger H, Boecker F, Packeisen J, Agelopoulos K, Poos K, Nadler W, Korsching E.
Analyzing the basic principles of tissue microarray data measuring the cooperative phenomena of marker proteins in invasive breast cancer.
OAB, 2013, 5(1):1-21 or arXiv, code on request.
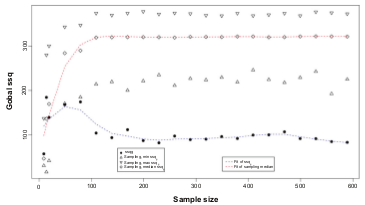

Hungermann D, Schmidt H, Natrajan R, Tidow N, Poos K, Reis-Filho JS, Brandt B, Buerger H, Korsching E.
Influence of whole arm loss of chromosome 16q on gene expression patterns in oestrogen receptor-positive, invasive breast cancer.
J Pathol. 2011 Aug;224(4):517-28.
Download
JP_supplementary_data.zip (14.2 MB)
Korsching E, Agelopolous K, Schmidt H, Buchroth I, Gosheger G, Wulfing P, Boecker W, Brandt B, Buerger H.
Improvements in the analysis strategy make single nucleotide polymorphism analysis a powerful tool in the detection and characterization of amplified chromosomal regions in human tumors.
Pathobiology. 2006;73(1):18-25.
Download
SNPtool.zip (4.3 MB)